BGSU RNA site Help
RNA 3D Motif Atlas
RNA 3D Motif Atlas is a comprehensive and representative collection of internal and hairpin loop RNA 3D motifs extracted from the Representative set (non-redundant lists) of RNA 3D structures. It is automatically updated every 4 weeks.
Display of Motif Atlas releases
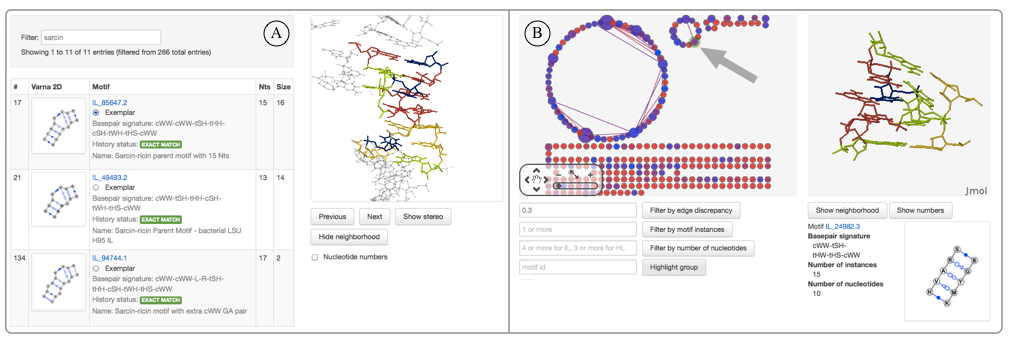
The list view (left) can be used to filter the list of motifs by any text appearing on the page. For example, one could quickly search for all sarcin-like motifs by typing “sarcin” in the filter box. The columns can be sorted by clicking on the table headers. The 3D structure displayed in Jmol is that of the exemplar motif instance (the first instance shown on the individual motif group pages).
The graph view (right) can be used for exploring structural similarities of motif groups. The motif groups are represented as nodes connected by edges if at least one instance from one motif matched at least one instance in another. Only those matches that met all quality assurance criteria are displayed. Clicking on each node displays its exemplar in 3D and some summary information. The users can filter the edges by the geometric discrepancy threshold value between the linked instances. Clicking on the edge retrieves the information about the connection between the two motifs and allows the user to compare the two motifs using the interface described above. The graph is produced by the Cytoscape Web browser plugin.
The graph view can also be used as a clustering validation tool. In general, in a good clustering only a small number of motifs should be connected at low discrepancy levels (<0.2-0.3 Å/nucleotide). When two motif groups are connected at very low discrepancies, then it is possible that one of the connected instances has been misclassified; such problems can be examined using the motif group comparison view described below.
Comparing Motif Atlas releases
Any pair of Motif Atlas releases can be compared. For example, let’s take a look at a comparison of two releases: 0.10 and 0.11.
Changes in PDB files
Every Motif Atlas release is based on a representative set (non-redundant list) of PDB files. When comparing two releases, it’s important to know how the input data changed. Representative sets (non-redundant lists) are quite stable in time, but PDB files from the older release can be replaced by newer PDB files or removed from the set altogether if the structure is obsoleted by PDB without replacement. PDB files from the newer release either replace representatives in the existing equivalence classes or come from new equivalence classes.
Changes in motif groups
It is helpful to track the motif groups with each release to distinguish new motifs from those that have been updated by adding, removing or replacing some motif instances. The use of a version number in motif group ids allows us to maintain the same base id for stable motif groups, while also recording when changes occur between releases. For example, the addition of new triple-sheared motif instances to the triple-sheared motif group need not result in the creation of a brand new id for the triple-sheared motif, because it already contains a large number of members, so only the version number will be incremented (for example, motif id IL_87904.1 has been incremented to IL_87904.2 and then to IL_87904.3).
Sankey diagrams
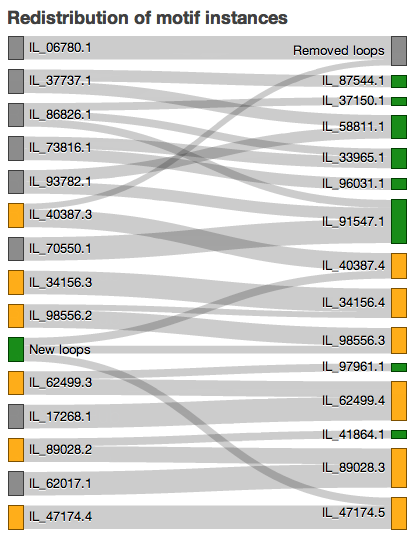
The Sankey diagrams show redistribution of motif instances (loops) between releases. The rectangles represent the motif groups and are colored as follows:
Added motif groups: green
Removed motif groups: grey
Updated motif groups: orange
Added motif instances: green
Removed motif instances: grey
The links between the releases represent motif instances that moved between different motif groups. You can hover over the links connecting the motif groups to see which motif instances were exchanged.
The diagrams are made with the Sankey d3 plugin.
Display of individual motif groups
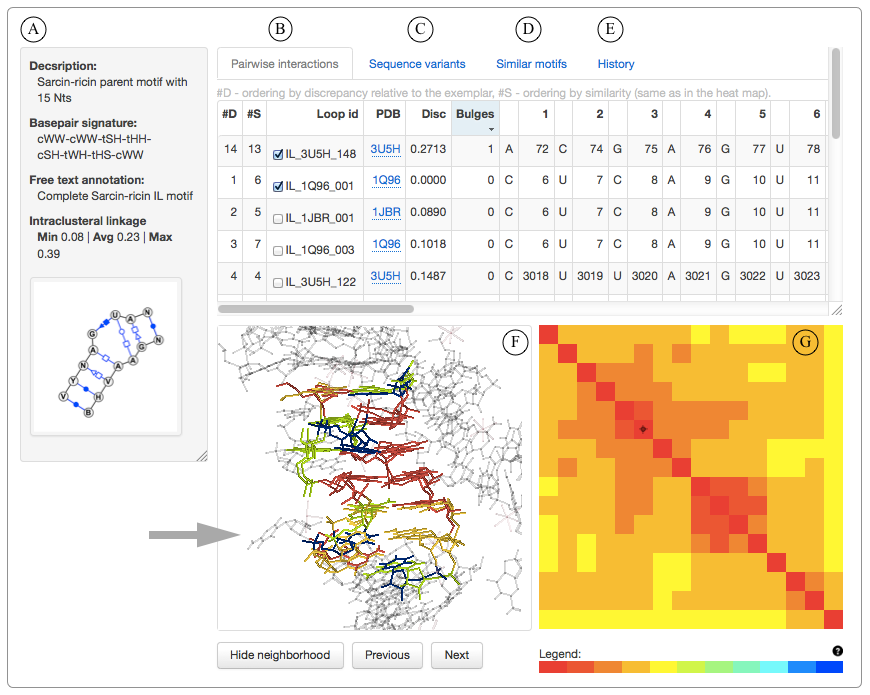
A) VARNA 2D diagram and manually curated description and free text annotation of the motif group. The annotations are automatically propagated to the successor motif groups as the Motif Atlas is updated.
B) The motif instances and the pairwise interactions between their constituent nucleotides are presented in a tabular form. The first instance in the list is the geometric centroid of the group and is the representative instance or exemplar. Other motif instances are ordered by their geometric discrepancy from the centroid (column #D). The instances can also be interactively sorted by similarity, which puts similar instances next to each other (column #S).
C) Sequence variability of the motif instances forming the motif group can be viewed for each motif.
D) The information about pairwise similarity between all motif instances obtained during all-against-all geometric FR3D comparisons is used to identify and rank geometrically similar motifs.
E) The releases in which the motif group is present are listed on the History tab. It also contains the detailed information about preceding and successor motif groups.
F) Jmol visualization of the motif instances. The user can view the 16A neighborhood to see bulged bases (shown with an arrow).
G) The similarity ordering, when applied to the matrix containing geometric discrepancies between all motif instance pairs, produces a heat map indicating similarity between each pair of instances in the motif group. Low geometric discrepancy (red and yellow colors) indicates high structural similarity while cyan and blue colors represent high geometric discrepancy between the structures. The heat map is interactive: clicking a cell above the diagonal shows 3D superposition of the two motif instances in the Jmol applet. Clicking a cell on the diagonal displays the corresponding motif instance, and clicking on a cell (i,j) below the diagonal superposes all motif instances from i to j and is equivalent to clicking all diagonal cells in the interval from (i,i) to (j,j). This functionality allows for intuitive examination of outliers and discovery of submotifs within the motif group.
Comparing motif groups
Any two motif groups, including those belonging to different Motif Atlas releases, can be compared using the interface shown in the figure below. The interface is available from the “Similar motifs” tab of the individual motif group pages as well as from the Motif Atlas release graph view.
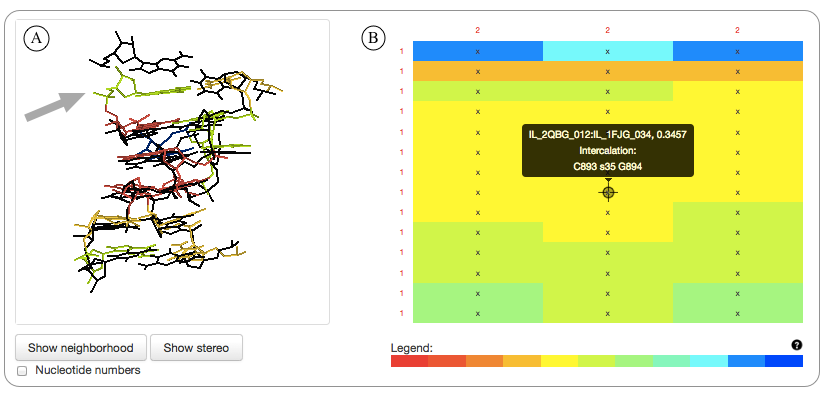
Web interface for comparing two motif groups. A) Jmol superposition of two motif instances belonging to different motif groups. The instances are separated in different groups because one of the instances has an intercalated nucleotide (location indicated by the grey arrow). B) The heatmap showing geometric discrepancy between 14 instances belonging to the first motif group (rows) and the 3 motif instances belonging to the second motif group (columns). Clicking on the cells brings up the 3D superposition of the corresponding motif instances in the Jmol window on the left. Disqualified pairs of instances are marked with an “x”. Detailed explanations for all disqualification decisions are also provided.
Polymorphic motifs
In the Motif Atlas there is a special view for identifying motifs with the same sequence (polymorphic motifs). This view is available from the release pages.
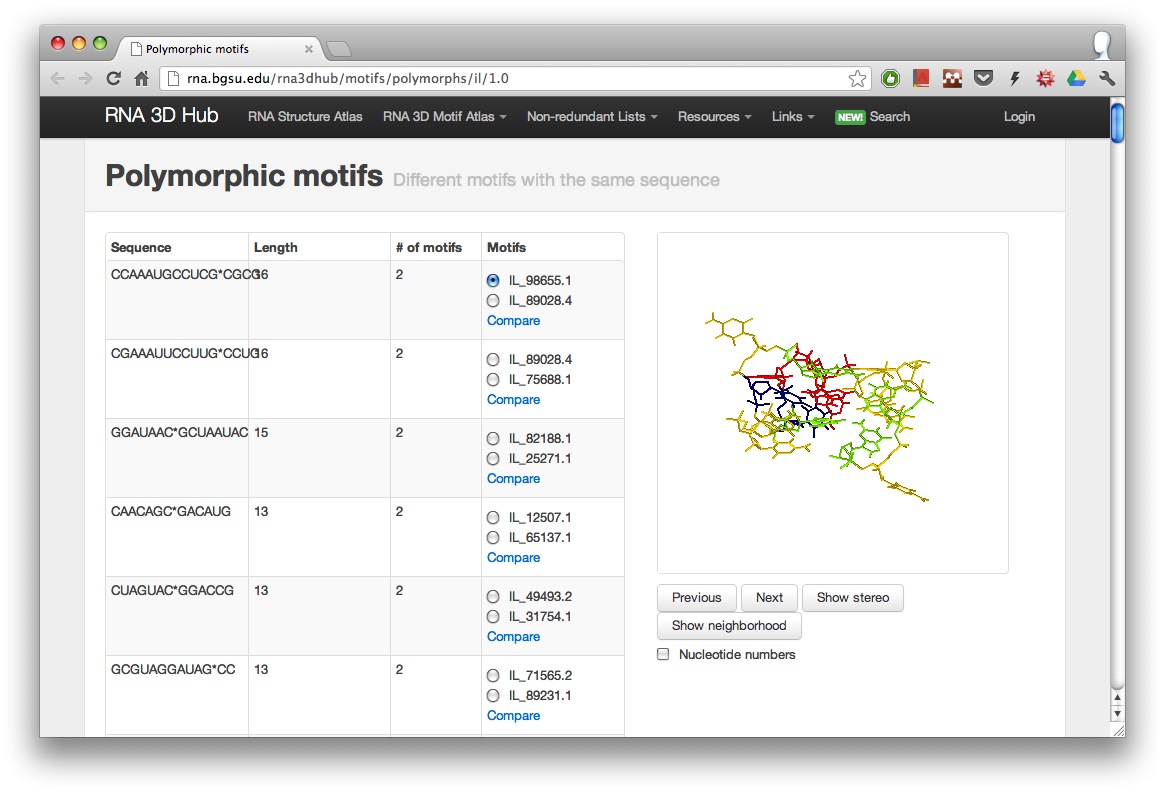
One can click on the compare link to study any motif pair. When two instances have identical sequences, the corresponding cell in the mutual discrepancy table will rock. For example, on this page a black cell is rocking. The cell is colored black if two instances didn’t match during pairwise all-against-all searches. When comparing the two instances in question (IL_2OE5_002 and IL_2FQN_002), it is clear that they belong to two different because motif groups IL_57785.5 and IL_54965.1 because they represent the decoding site in two states.
Browsing ribosomal secondary structures
Secondary structures provide a convenient way of exploring ribosomal RNAs and are familiar to many scientists. To facilitate navigating RNA 3D Motif Atlas, we labeled publicly available 2D diagrams of rRNAs with links to RNA 3D motif instances from the Motif Atlas, which can be accessed at http://rna.bgsu.edu/rna3dhub/motifs/2ds.
Currently we provide E.coli (SSU and LSU), T.thermophilus (SSU and LSU), H. marismortui (LSU), D. radiodurans (LSU), S.cerevisiae (LSU). Here is a 2D with motif annotations from E. coli 16S:
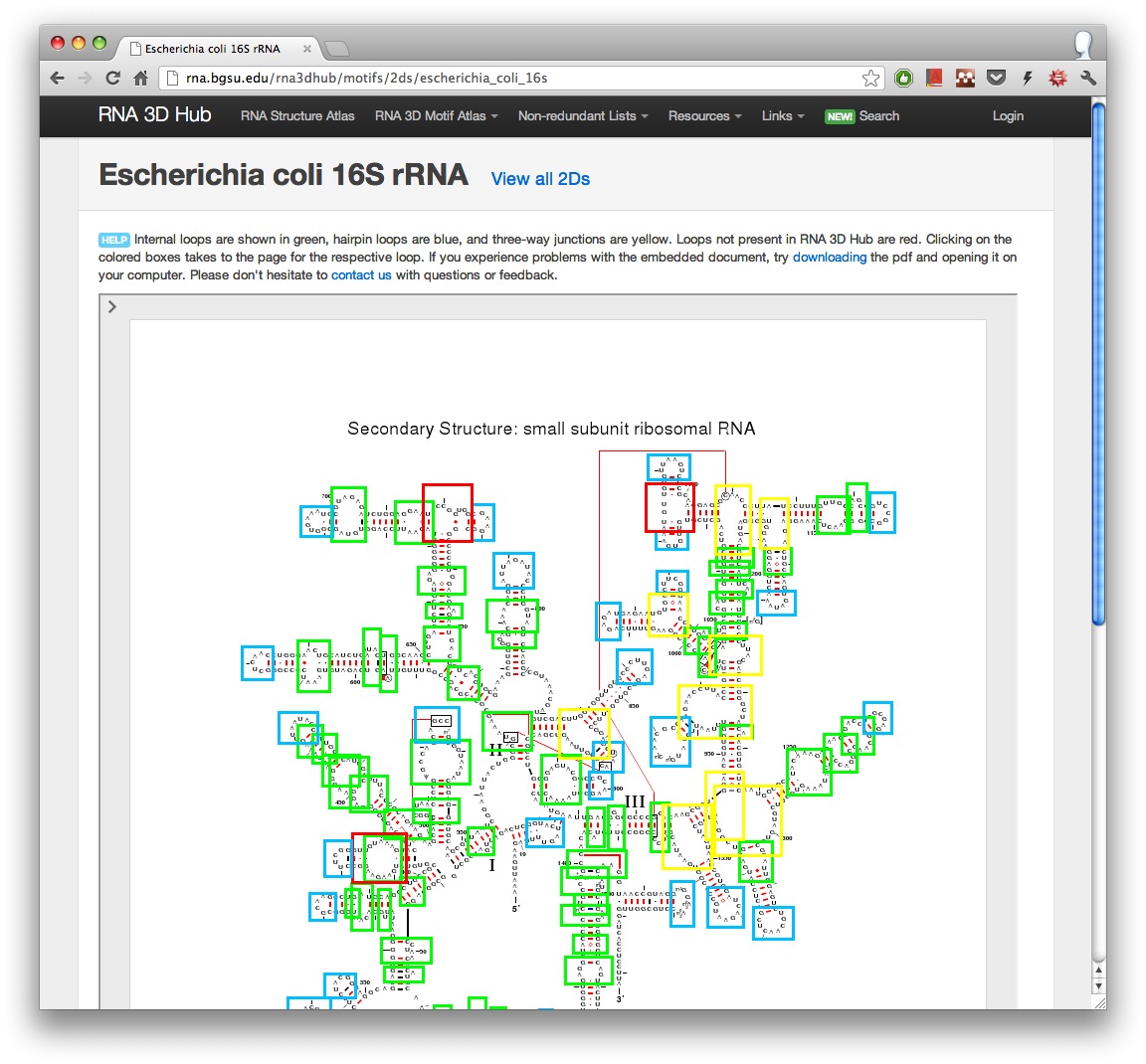
Internal loops are shown in green, hairpin loops are blue, and three-way junctions are yellow. Loops not present in our database are red.
To navigate to a motif group, first click on one of the colored boxes. You will be taken to the page for the respective loop. At the bottom of the page you will find the link to the motif entry in the Motif Atlas. For example, at the time of writing internal loop IL_2AW7_065 was assigned to motif IL_13959.3. Clicking on that link will show other motif instances assigned to the same motif group.
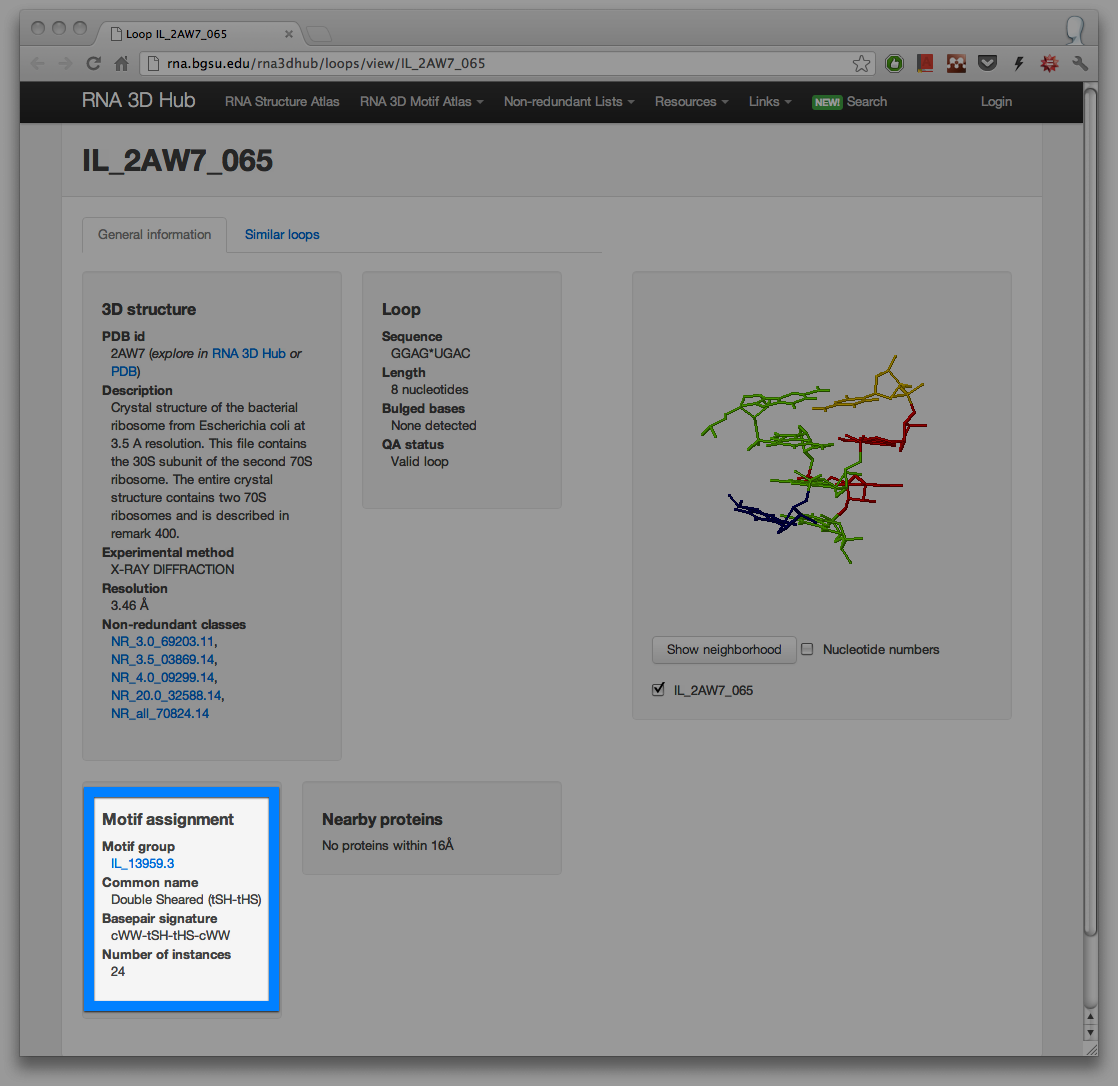
We are also working on a new way of browsing RNA secondary structures dynamically annotated with data from our database. For preview, take a look at the annotated secondary structure diagram from 16S E. coli: http://rna.bgsu.edu/rna3dhub/pdb/2AW7/2d.
Searching Motif Atlas
Currently RNA 3D Motif Atlas can be searched in three ways. Firstly, the Motif Atlas is integrated with a customized Google search engine, which provides a familiar and flexible way of searching across the entire RNA 3D Motif Atlas by PDB id, motif instance sequence (in 5’ to 3’ direction), or any other text appearing on the Motif Atlas web pages.
Secondly, to search within a single Motif Atlas release, the users can efficiently filter the motif collection by the motif size or by common name on the motif release pages where it is also possible to explore motif groups visually using Varna-generated 2D diagrams of motif exemplars, which can also be viewed in 3D using Jmol.
Finally, several ribosomal secondary structure diagrams of different organisms have been annotated with loop ids. The users can click on the 2D diagrams, which are linked to the Motif Atlas pages with the most recent motif assignments and other annotations. These diagrams can be used to compare RNA 3D motifs found in homologous locations in different organism.
Downloading data
Motif group data are provided for download in csv and json formats. The csv files only contains the aligned nucleotides. The json files also include basepair signatures, common names and other useful information in addition to nucleotide-to-nucleotide alignments of motif instances.
For any motif group or Motif Atlas release, the users can either download motif descriptions manually using the online interface or by constructing and accessing the URLs programmatically. A stable URL linking to the most current Motif Atlas release is also provided, which can be used for integration with other resources. Detailed instructions are also available.
To describe nucleotide alignments, we use a stable nucleotide naming scheme, which uniquely and unambiguously identifies any residue in the PDB database. Each nucleotide is assigned a string identifier containing pdb id, model number, chain id, component id, alternate id, insertion code, and symmetry operation used to obtain the coordinates of the nucleotide. The nomenclature is going to be presented in more detail elsewhere. Please refer to the documentation describing the format.
Implementation
The RNA 3D Motif Atlas data analysis pipeline is fully automated and runs every four weeks. The all-against-all structural alignments and geometric discrepancy calculations are carried out using the FR3D software suite and are performed in Matlab under control of Python scripts using Mlabwrap. The data are imported into a MySQL database, which serves as the backend for the Motif Atlas. The site is implemented in PHP and Javascript using open-source industry-standard frameworks including CodeIgniter, jQuery,SQLAlchemy, and Twitter Bootstrap. In addition, Jmol is used for 3D structure and Cytoscape Web for graph visualization. All software developed as part of this project can be found on GitHub.
Using Jmol Applet for viewing Motif Atlas
As HTML5 becomes the mainstream web technology, Java applets are no longer available on some platforms due to security issues. In the future, the Motif Atlas will switch to a pure javascript implementation of Jmol, such as jsmol. However, for now some users may need to experiment with their browser(s) to get Jmol working.
It is difficult to describe an ideal environment for running Jmol applets because Jmol availability depends on many factors, such as the operating system, the browser, and the version of Java. Sometimes a computer can have several versions of Java installed in parallel, and it’s important to make sure that the browser is configured to use the right version.
To make matters worse, the same computer that runs Jmol one day may not be able to run it later. For example, in 2013 Apple several times remotely disabled Java on Macs running the latest Mac OS 10.8, which turned off all Java plugins, including Jmol. The users had to manually update their Java and configure their browsers to get Jmol working again.
Unfortunately, we cannot provide up-to-date information about Jmol availability. Users are encouraged to use their favorite search engines to diagnose the problem and contact us if additional assistance is desired.
Representative sets (non-redundant Lists)
Downloading data
It’s possible to download the lists in csv format using the web interface:

The csv files are formatted as follows:
"Equivalence class id","Representative structure","Comma-separated list of the equivalence class members"
For example, http://rna.bgsu.edu/rna3dhub/nrlist/download/0.87/1.5A/csv:
“NR_1.5_23181.1″,”2ASB”,”2ASB”
“NR_1.5_11995.1″,”3HGA”,”3HGA”
“NR_1.5_76588.1″,”397D”,”397D”
“NR_1.5_49442.1″,”2R22″,”2R22″
“NR_1.5_27899.1″,”2Y8Y”,”2Y8Y”
“NR_1.5_93883.1″,”3ND3″,”3ND3,3ND4″
Also, the data can be accessed automatically using wget or any other program. For example, here is a sample Python script for downloading the current non-redundant lists: https://gist.github.com/3761913
The detailed instructions for downloading the date are available.
RNA Structure Atlas
The RNA Structure Atlas provides a new way of browsing our database by PDB id. It integrates the information from our databse and presents it in a streamlined interface, which can help explore the structural database in a more efficient way.
RNA Structure Atlas uses Chosen, an excellent javascript plugin, which powers the navigation dropdown list. We also use the JmolTools plugin for dynamic retrieval and visualization of 3D structural fragments in Jmol. Please feel free to get in touch with any feedback.
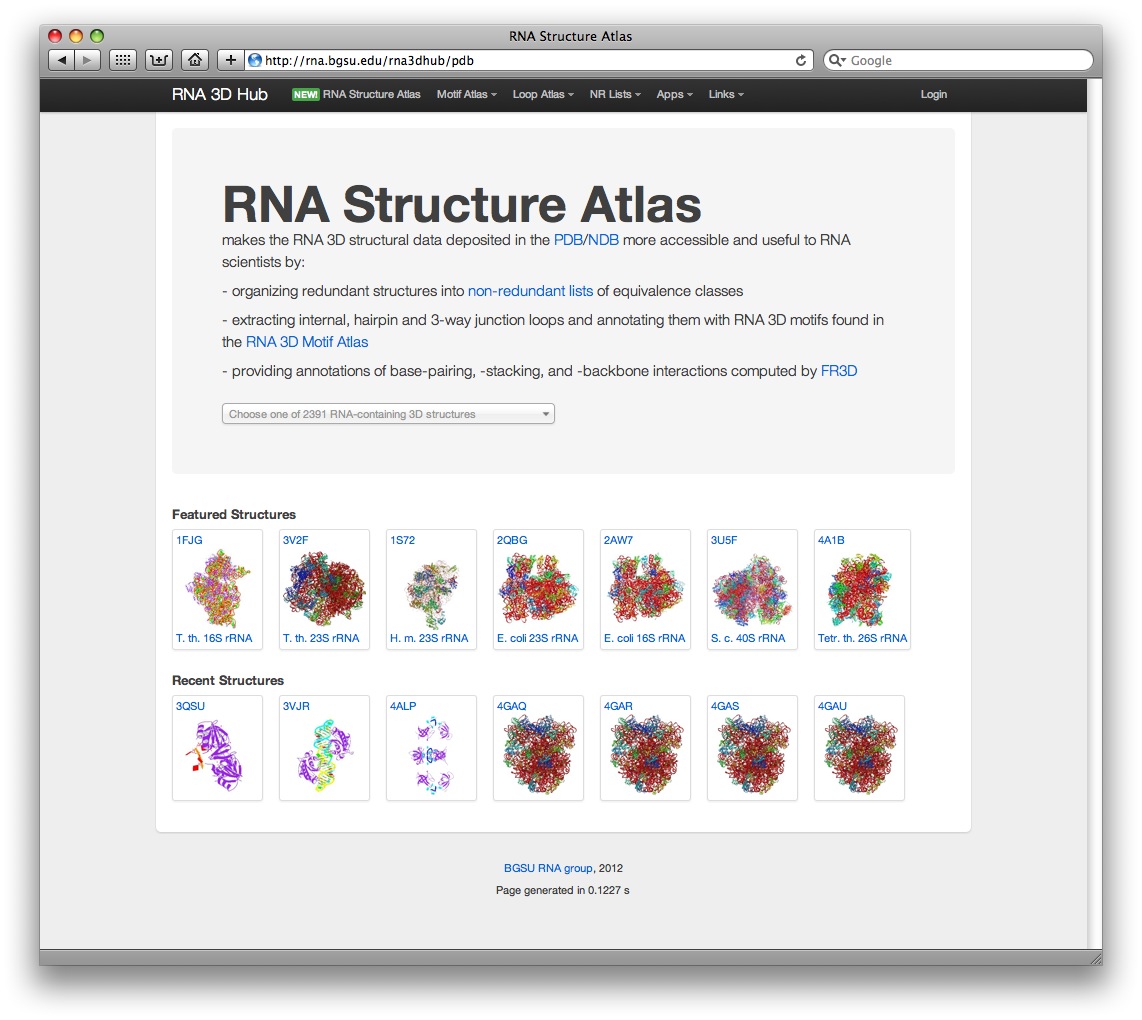
Individual structure pages
Each PDB file has its own webpage, for example, here is a summary page for PDB 1S72:
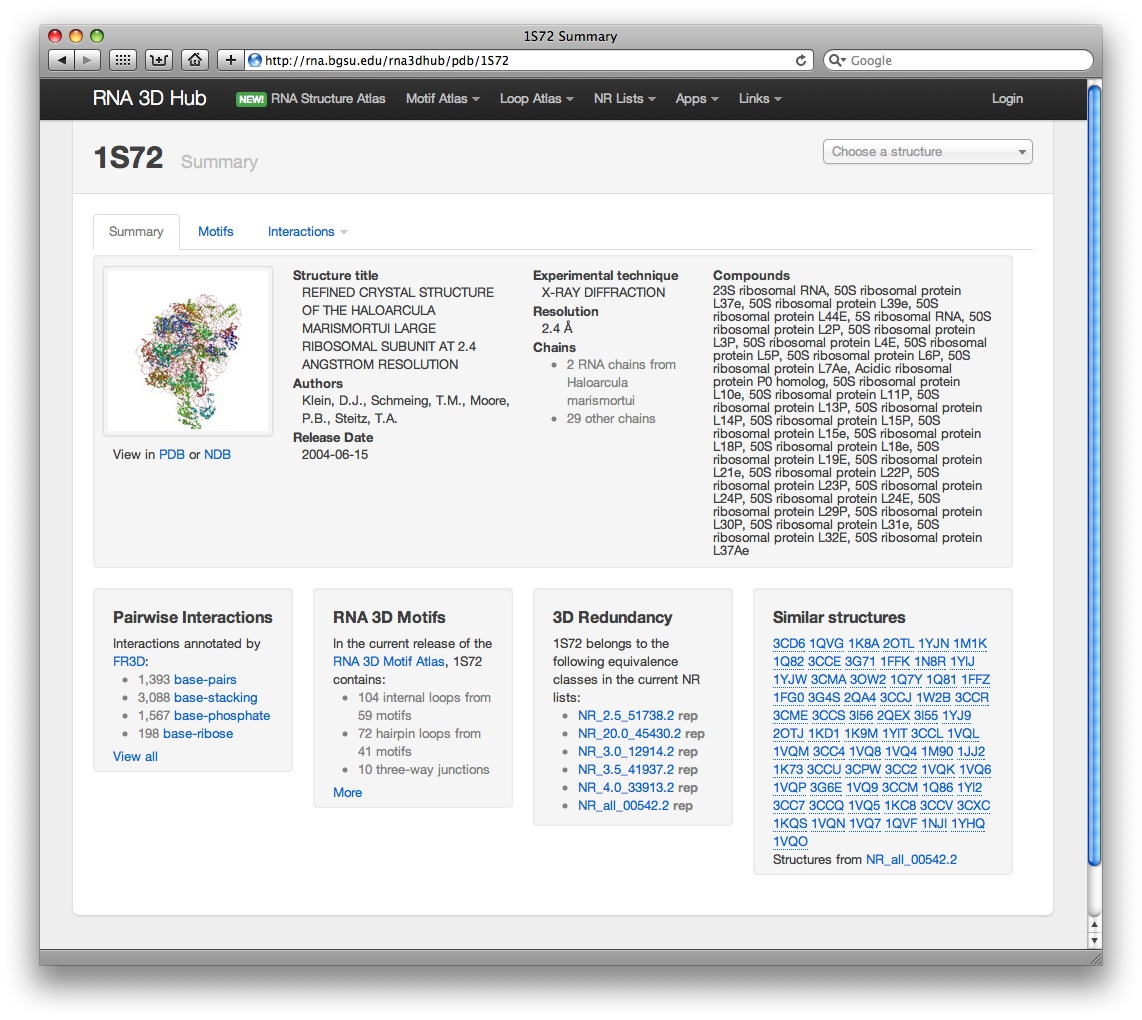
From this page the users can navigate to other webpages where they can:
- view and download pairwise interaction annotations produced by FR3D, including base-pairing, base-stacking, and base-phosphate interactions
- view all internal, hairpin and junction loops found in the 3D structure
- identify similar RNA 3D structures from the representative set (non-redundant lists)
Pairwise interactions pages
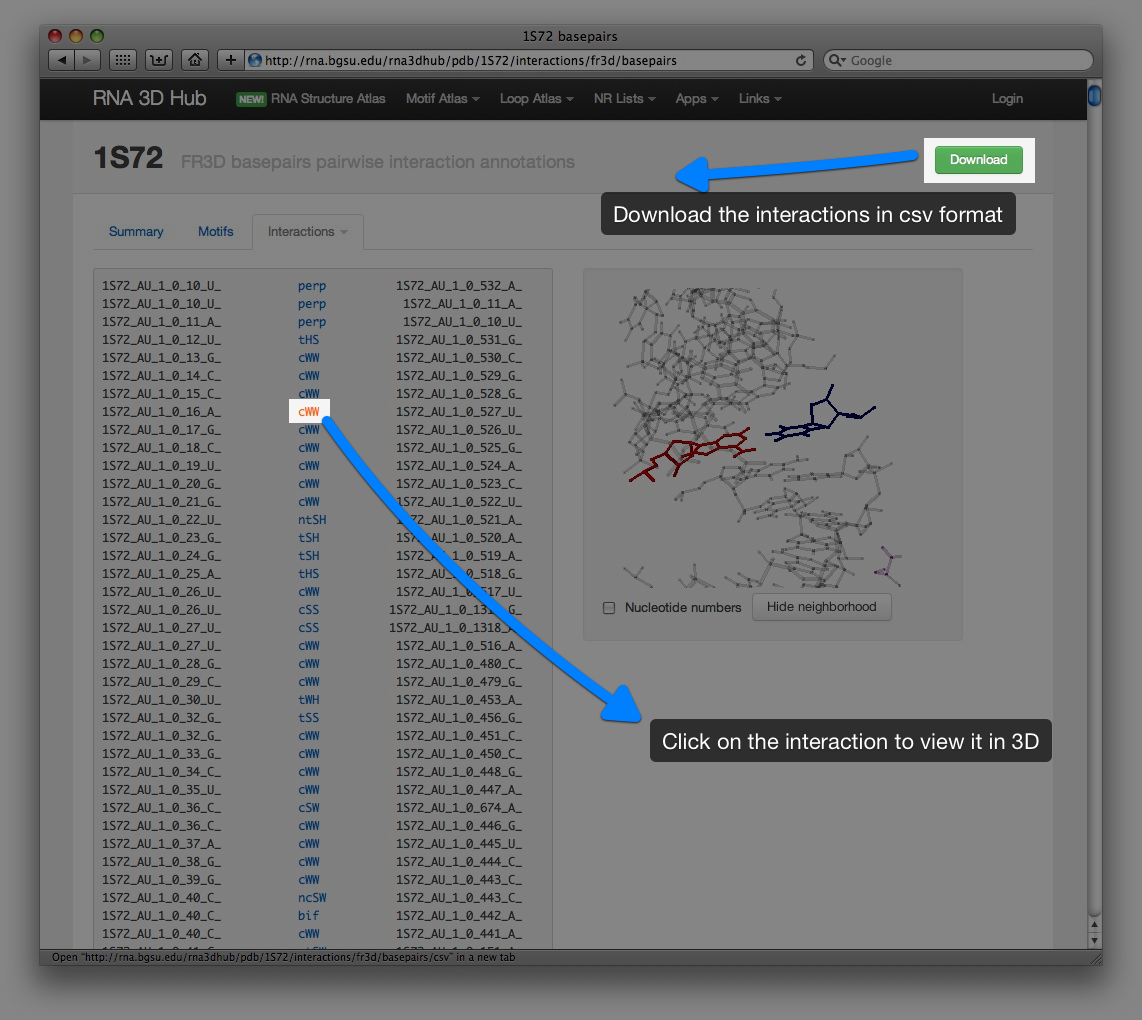
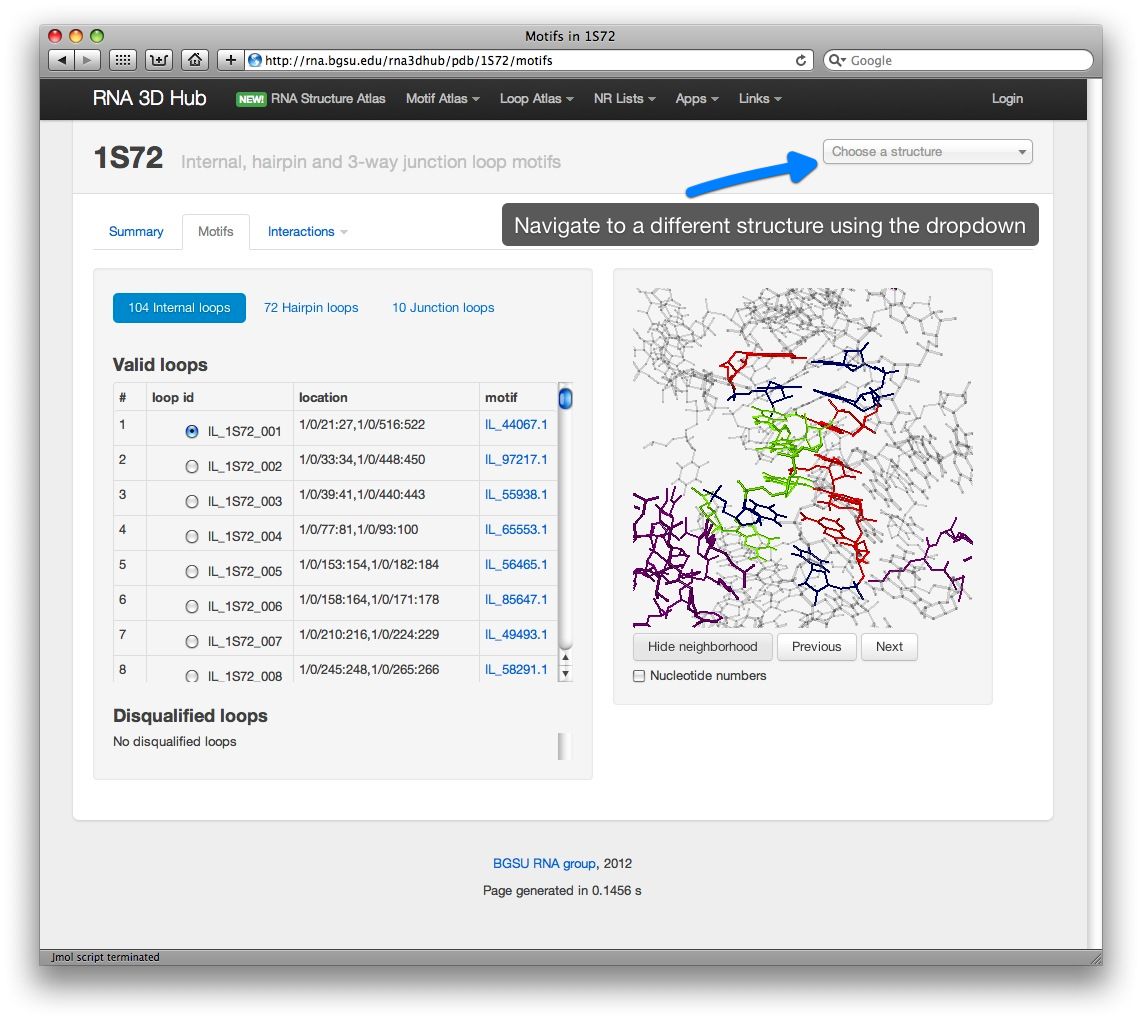
Motif pages
Some structures are also annotated with recurrent motifs from the RNA 3D Motif Atlas. In the future, all structures will be annotated with motifs.
Updated: 08/01/2025 05:42PM